AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
Back to Blog
Graphpad prism heatmap12/6/2023  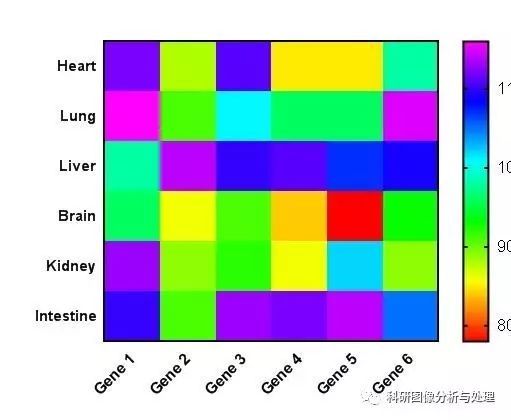
Sample tags allow working one step above FCS file names, using the variables present in the experiment to create meaningful figures.  
Unstimulated, Stimulated) and individuals (HD1, HD2, HD3, HD4). Sample tags were assigned to each FCS file in order to define conditions (e.g. A viSNE run was performed within the Cytobank platform on all files, in order to obtain the same map across all samples. In our example, we had eight FCS files, derived from four different healthy donors, each one either stimulated or not with anti-CD3/CD28 beads. treatments) as they may result in the presence or absence of a specific island in the viSNE map. Indeed, with viSNE maps 1 you can quickly identify and visualize changes among your samples due to different experimental conditions (i.e. This makes viSNE a good option when you are exploring a way that can help define biological effects in a fast and visual fashion. 
In a viSNE map, cells that are phenotypically similar will be close to each other and may form an island. With advances in technologies, the number of parameters that can be analyzed together in each flow cytometry panel keeps increasing, and if we sum this up with the need for testing and analyzing many samples together, the analysis step can become overwhelming.ĭimensionality reduction tools, such as viSNE maps 1, enable you to reduce high dimensional data into two dimensions, thereby enabling rapid exploratory analysis and visualization of complex results. Thus, many samples that underwent the same treatment need to be visualized. When looking at how a specific experimental condition, i.e., a specific treatment, can affect cell populations present in your samples, you need to be sure that the effect you’re seeing is the consequence of a common biological phenomenon, not specific for one single sample. Transformed data was exported using the Kaluza Cytobank Plugin. Data were acquired on a 13 detector / 3 laser CytoFLEX V5-B5-R3 (PN: C09734) compensation and logicle transformation were adjusted using Kaluza Analysis software. Lineage markers and cytokine expression were assessed after six hours with a 13-color antibody cocktail described below (Table 1). To generate the data used in this Application Note, cryopreserved human peripheral blood mononuclear cells (PBMC) from healthy donors were thawed and rested overnight for cell function recovery stimulated using Dynabead T-cell Activator anti-CD3/CD28 (ThermoFisher Scientific) in presence of protein transport inhibitor (Brefeldin A). How to use heatmaps to help you identify the phenotype of a specific cluster/metacluster.How FlowSOM can help you in highlighting differences in abundance of specific clusters in your experimental groups.How to leverage a dimensionality reduction algorithm like viSNE for visual comparisons between groups of samples.Liquid Handling and Scheduling Software.CytExpert Software for the CytoFLEX Platform.Plate Loader Options for the CytoFLEX Platform.CytoFLEX Violet-Blue-Red Series Upgrades.Labware for Liquid Handling Instruments.Biomek NGeniuS Reaction Vessel, 24 Well, 64/Box.Lysing, Fixative, and Permeabilizating Reagents.RESOURCE Contract Manufacturing Services.HIAC 9703+ Pharmaceutical Particle Counter.Biomek NGeniuS Next Generation Library Prep System.Cell Counters, Sizers and Media Analyzers.Air Particle Counters for Cleanroom and Environmental Monitoring. 
0 Comments
Read More
Leave a Reply. |
 RSS Feed
RSS Feed
